Author Template - 2. Code
Abstract¶
Contains examples of all MyST markdown code syntax
Code Section¶
This chapter demonstrates advanced usage of {code-cell}.
All the examples are themed around MOF structure modeling / adsorption simulation / materials property calculation.
1. Static Code (Not Executed)¶
Block content:
:::{code} python
def mof_density(M, V):
return M / V
print(mof_density(1000, 500))
:::def mof_density(M, V):
return M / V
print(mof_density(1000, 500))Suitable for: Concept explanations / example code snippets
2. Executable code-cell¶
(Requires kernelspec at the top of the file and running myst start --execute)
:::{code-cell} python
import numpy as np
print("Example: generate random porosity:", np.random.rand())
:::import numpy as np
print("Example: generate random porosity:", np.random.rand())Example: generate random porosity: 0.8708862896410129
3. Add Metadata¶
Metadata can control the way code is executed and displayed.
3.1 Hide Input (only show output)¶
Block content:
:::{code-cell} python
:tags: hide-input
print('MOF is great')
:::Source
print('MOF is great')MOF is great
3.2 Hide Output (only show code)¶
Block content:
:::{code-cell} python
:tags: hide-output
x = 42
x
:::x = 42
xOutput
423.3 Completely Remove Input (code is invisible in the final webpage)¶
Block content:
:::{code-cell} python
:tags: remove-input
print("This code is not visible in the final webpage, but still executed")
:::This code is not visible in the final webpage, but still executed
3.4 Completely Remove Output (output not visible on webpage)¶
Block content:
:::{code-cell} python
:tags: remove-output
print("This code is not visible in the final webpage, but still executed")print("This code is not visible in the final webpage, but still executed")Frequently Used Tags¶
| tag | Meaning |
|---|---|
| hide-input | Hide code, show output only |
| hide-output | Hide output |
| remove-input | Completely remove input on the page |
| remove-output | Completely remove output |
| raises | Specify expected exceptions |
| cache | Allow execution caching |
4. Plotting¶
The following example shows a plot of the relationship between MOF porosity and surface area.
import numpy as np
import matplotlib.pyplot as plt
phi = np.linspace(0.1, 0.9, 9)
sa = 1500 * phi * 2
plt.figure(figsize=(6,4))
plt.plot(phi, sa, marker="o")
plt.xlabel("Porosity φ")
plt.ylabel("Surface Area (m²/g)")
plt.title("MOF Porosity vs Surface Area (illustration)")
plt.grid(True)
plt.show()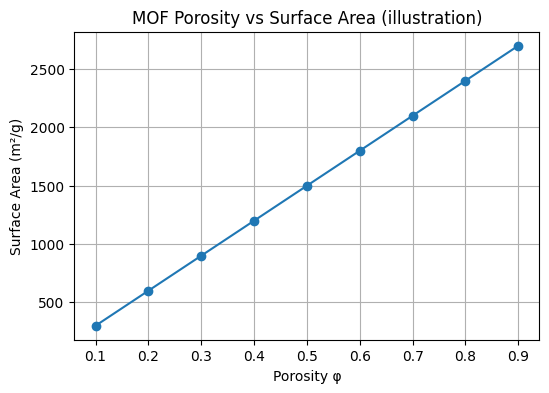
5. Interactive Visualization¶
The following example uses Altair to create a simple interactive visualization of MOF porosity and surface area. You can brush-select a data range to highlight selected points and view corresponding statistics.
Source
import altair as alt
import numpy as np
import pandas as pd
# Generate MOF data with different types
np.random.seed(42) # For reproducibility
mof_types = ['MOF-5', 'UiO-66', 'HKUST-1', 'MIL-101(Cr)', 'NU-1000']
phi_range = np.linspace(0.1, 0.9, 25)
# Create data for different MOF types with varying relationships
data_list = []
for mof in mof_types:
# Different scaling factors for different MOFs
if mof == 'MOF-5':
scale_factor = 2.0
base_sa = 1500
elif mof == 'UiO-66':
scale_factor = 1.8
base_sa = 1200
elif mof == 'HKUST-1':
scale_factor = 2.2
base_sa = 1800
elif mof == 'MIL-101(Cr)':
scale_factor = 2.5
base_sa = 2000
else: # NU-1000
scale_factor = 2.3
base_sa = 1900
for phi in phi_range:
# Surface area relationship: varies by MOF type
sa = base_sa + scale_factor * 5000 * phi
data_list.append({
'MOF_Type': mof,
'Porosity': round(phi, 2),
'Surface_Area': round(sa, 1)
})
df = pd.DataFrame(data_list)
# Brush selection for interactive filtering
brush = alt.selection_interval(encodings=['x', 'y'])
# Main scatter plot with lines
points = alt.Chart(df).mark_point().encode(
x=alt.X('Porosity:Q',
axis=alt.Axis(title='Porosity φ', format='.2f'),
scale=alt.Scale(domain=[0.1, 0.9])),
y=alt.Y('Surface_Area:Q',
axis=alt.Axis(title='Surface Area (m²/g)'),
scale=alt.Scale(domain=[0, 15000])),
color=alt.condition(brush, 'MOF_Type:N', alt.value('lightgray')),
#size=alt.condition(brush, alt.value(80), alt.value(40)),
tooltip=[
alt.Tooltip('MOF_Type:N', title='MOF Type'),
alt.Tooltip('Porosity:Q', title='Porosity', format='.2f'),
alt.Tooltip('Surface_Area:Q', title='Surface Area', format='.1f')
]
).add_params(brush).properties(
width=600,
height=400,
title='MOF Porosity vs Surface Area'
)
# Bar chart showing count of selected MOF types
bars = alt.Chart(df).mark_bar().encode(
y=alt.Y('MOF_Type:N', axis=alt.Axis(title='MOF Type')),
color='MOF_Type:N',
x=alt.X('count():Q', axis=alt.Axis(title='Count'))
).transform_filter(brush).properties(
width=600,
height=200,
title='Selected MOF Types Distribution'
)
# Combine charts vertically
points & barsInteractive Functionality Instructions:
Brushing Selection: Drag the mouse on the main chart to select a data region; selected points will be highlighted (in color), unselected will be gray
Statistics Chart: The bar chart below will show the count distribution of each MOF type in the selected region in real time
Tooltip: Hover over any data point to see details
6. Pandas Table¶
import pandas as pd
df = pd.DataFrame({
"MOF": ["MOF-5", "UiO-66", "HKUST-1"],
"Metal": ["Zn²⁺", "Zr⁴⁺", "Cu²⁺"],
"Surface Area (m²/g)": [3800, 1200, 1500]
})
df7. Structure Visualization (ASE)¶
(If your kernel has ase installed)
7.1 Build molecule via ASE and show interactively¶
from ase.build import molecule
from ase.visualize import view
try:
# Build a simple molecule with ASE
water = molecule('H2O')
# Visualize the molecule
disp_water = view(water, viewer='x3d')
display(disp_water)
except:
print('ASE is not installed, so the example is skipped')7.2 Read and show structure from local file (CIF / POSCAR etc.)¶
from ase.io import read
# Read structure from a local file, e.g. CIF/PDB
mof_from_file = read("../data/MOF-5.pdb")
# View the periodic structure
disp_mof=view(mof_from_file, viewer='x3d')
display(disp_mof)