MOF Book Example Chapter 1 (Basics)
This chapter introduces the basic concepts of MOFs and demonstrates fundamental features and formatting.
Chapter 1: Basic Concepts and Structure of MOFs¶
1.1 What is a MOF?¶
A Metal–Organic Framework (MOF) is a class of porous crystalline material constructed from metal nodes and organic linkers.
Metal nodes: Typically metal ions or clusters (such as Zn²⁺, Cu²⁺, Zr⁴⁺, etc.)
Organic linkers: Multidentate ligands containing carboxyl, pyridyl, or other coordinating groups
Features: Exceptionally high specific surface area, tunable structure, and versatile functionalization
The main characteristics of MOF materials are their extremely high specific surface area and tunable pore structures. You can think of them as nano-sized Lego blocks.
As shown in the literature Yaghi, 2025[1]
1.2 Typical Structure Illustration and Unit Cell¶
1.2.1 Structure Illustration¶
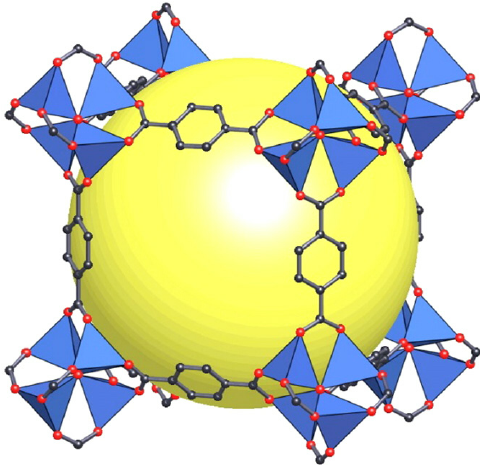
Figure 1:A schematic diagram of a MOF structure.
1.3 Simple Crystallographic Expressions¶
Below are formulas describing the basic cell size and porosity of a MOF framework.
1.3.1 Volume and Porosity¶
Unit cell volume:
If the void volume of a MOF, , is obtained via molecular simulation or adsorption experiments, the porosity can be expressed as:
1.4 Table: Some Typical MOFs¶
| Name | Metal Center | Organic Linker | Features |
|---|---|---|---|
| MOF-5 | Zn²⁺ | 1,4-benzenedicarboxylate (BDC) | Early classic MOF, high surface area |
| UiO-66 | Zr⁴⁺ | BDC | Excellent thermal/chemical stability |
| HKUST-1 | Cu²⁺ | 1,3,5-benzenetricarboxylate (BTC) | Classic copper-based MOF |
| MIL-101(Cr) | Cr³⁺ | BDC | Extra-large pore volume, good water stability |
1.5 Describing MOFs with Python Code¶
def describe_mof(name, metal, linker):
return f"{name} is a MOF with {metal} as the metal center and {linker} as the organic linker."
print(describe_mof("UiO-66", "Zr⁴⁺", "BDC"))The above is a Python code example for MOF description, using the describe_mof function.
1.6 Calculating the Relationship Between Porosity and Pore Volume in Python¶
Source
import math
def pore_volume(cube_length_nm, void_fraction):
"""
Simple cubic model.
cube_length_nm: cell edge length (nm)
void_fraction: porosity (void fraction)
Returns pore volume (nm^3)
"""
v_cell = cube_length_nm**3
return v_cell * void_fraction
for phi in [0.3, 0.5, 0.8]:
print(f"Porosity {phi:.1f} -> Pore volume: {pore_volume(2.5, phi):.2f} nm^3")1.7 Simple 2D Plot Example¶
import numpy as np
import matplotlib.pyplot as plt
phis = np.linspace(0.1, 0.9, 9)
surface_area = 1500 * phis * 2 # Hypothetical relation
plt.figure()
plt.plot(phis, surface_area, marker="o")
plt.xlabel("Porosity φ")
plt.ylabel("Surface Area (m²/g)")
plt.title("Simple Relationship Between Porosity and Surface Area")
plt.grid(True)
plt.show()1.8 An interactive MOF structure¶
Source
from ase.io import read
from ase.visualize import view
# Read structure from a local file, e.g. CIF/PDB
mof_from_file = read("../data/MOF-5.pdb")
# View the periodic structure
disp_mof=view(mof_from_file, viewer='x3d')
display(disp_mof)1.9 Exercise Zone (task list)¶
This is a review about AI for MOFs.
- Zheng, Z., Rampal, N., Inizan, T. J., Borgs, C., Chayes, J. T., & Yaghi, O. M. (2025). Large language models for reticular chemistry. Nature Reviews Materials, 10(5), 369–381. 10.1038/s41578-025-00772-8